Oncomedicine 2016; 1:25-27. doi:10.7150/oncm.16256 This volume Cite
Short Research Paper
ERCC1 Biomarker in Colorectal Cancer: To Induce or Not to Induce? This Is the Matter!
1. Medical Oncology Department, Università Cattolica del Sacro Cuore, Rome, Italy;
2. Department of Biochemistry, Università Cattolica del Sacro Cuore, Rome, Italy;
3. Department of Medical Oncology, Fondazione Poliambulanza, Brescia, Italy.
Received 2016-7-20; Accepted 2016-8-28; Published 2016-10-10
Abstract
In this short research communication, using circulating tumor cells (CTCs) as a “dynamic” tissue, we found that ERCC1 induction is significantly correlated with clinical outcome in colorectal cancer patients treated with oxaliplain-based therapy. This is the first report proposing the ability to induce ERCC1 as a blood-based biomarker for oxaliplatin resistance in metastatic colorectal cancer.
Keywords: CTC, ERCC1, oxaliplatin, colorectal cancer
Introduction
Oxaliplatin is a milestone of colorectal cancer therapy, but it is still lacking of a validated predictive biomarker of response (1). For many years ERCC1 have been studied as predictive factor of response to platinum compounds in many types of cancer (2). A lot of studies have investigated the ERCC1 mRNA and/or protein expression, on primary tumor in colorectal cancer, as oxaliplatin's predictor factor, but the translation into clinical practice has not yet occurred for contrasting result and the lack of prospective evidence (3). We recently demonstrated in vitro that KRAS mutated (mt) cell lines were more sensitive to oxaliplatin due to their inability to induce ERCC1 after drug exposure (4). Using circulating tumor cells (CTCs) as a “dynamic” tissue, in this study we aim to confirm in vivo the inducibility of ERCC1 gene expression and its relationship with clinical outcome in colorectal cancer patients treated with an oxaliplatin based therapy.
Materials and Methods
We collected peripheral blood samples (10 ml) from metastatic colorectal cancer patients treated with oxaliplatin-based first-line regimen at 0 and 48 hours during the first cycle of chemotherapy. CTC was detected by the AdnaTest ColonCancerSelect and the AdnaTest ColonCancerDetect (AdnaGen AG, Langenhagen, Germany) system followed by multiplex RT-PCR including ERCC1 transcript. In CTC-positive ERCC1-positive patients, ERCC1 mRNA expression was measured using a quantitative RT-PCR method. We evaluated the relationship between ERCC1-induction and KRAS mutational status and we correlated the capacity of ERCC1-induction with clinical outcome (progression free survival, PFS) using the Kaplan-Meier method, the differences were compared with the use of log-rank test considering statistically significant a p value of ≤0.05.
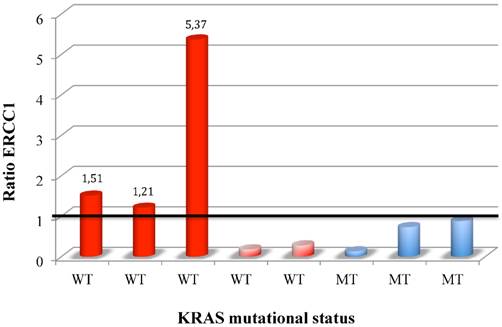
ERCC1 induction in CTC colorectal patients stratify for KRAS mutational status.
Progression free survival in CTC ERCC1+ve patients stratify for ERCC1 induction. median PFS of patients with ERCC1 induction was significantly shorter than that observed in patients with stable or reduced ERCC1 (2,7 months vs 8,8 months; HR: 0.2 IC95% 0.1-0.33, p 0.02).
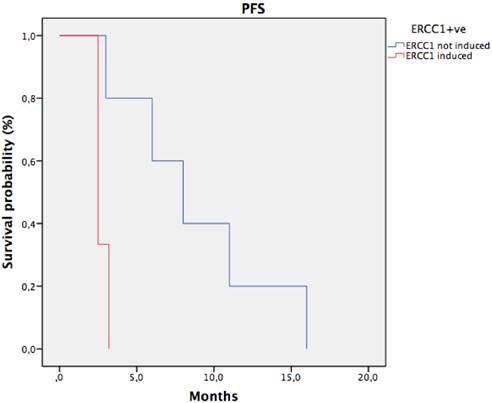
Results
38 patients were enrolled, 19 were KRAS wild-type (wt) and 19 KRAS mt. CTCs were detected in 12 patients (31,5 %). ERCC1 was expressed in 8 CTCs-positive (ERCC1-CTC+ve) patients (21%), 5 KRAS wt and 3 KRAS mt. After oxaliplatin exposure, among ERCC1-CTC+ve patients, only 3 showed a significant induction of ERCC1 expression (Figure 1); interestingly all of them were KRAS wt and experimented a rapid progression of disease. Notably median PFS of patients with ERCC1 induction was significantly shorter than that observed in patients with stable or reduced ERCC1 (2,7 months' vs 8,8 months; HR: 0.2 IC95% 0.1-0.33, p 0.02) (Figure 2).
Discussion
We found that ERCC1 induction is significantly correlated with clinical outcome in colorectal cancer patients treated with oxaliplain-based therapy. Interesting all patients that were unable to induce ERCC1 showed a significantly better outcome. According to our hypothesis of association between KRAS mutations and lack of ERCC1-inducibility, none of the mutated patient was able to induce ERCC1 after drug exposure and experienced a good prognosis.
Although based on a small sample size, this is the first report proposing the ability to induce ERCC1 as a blood-based biomarker for oxaliplatin resistance in colorectal cancer. Based on our hypothesis, a possible explanation for lack of correlation between basal ERCC1 tissue levels and response to oxaliplatin is that analysis on tumor biopsy before therapy gave only partial information losing the possibility to recognize the inducibility of ERCC1, one of the major hypothetic mechanism of drug resistance (5).
CTCs offer a unique opportunity as they are a vital surrogate for the tumor itself and could ensure not invasive liquid biopsies for serially monitoring dynamic processes as enzymatic induction. Our finding underlies the complex role of ERCC1 as a biomarker, has to be considered as a proof of concept, so exploratory in nature but need to be confirmed in larger studies because may have important clinical implications.
Author Contributions
Study concept and design: Di Salvatore, Orlandi, Paolillo.
Acquisition, analysis, or interpretation of data: Di Salvatore, Orlandi, Paolillo, Aroldi, Rodriquenz.
Drafting of the manuscript: Di Salvatore, Orlandi.
Critical revision of the manuscript for important intellectual content: Capolongo, Barone.
Statistical analysis: Orlandi.
Study supervision: Capolongo, Barone.
Conflict of Interest
The authors have completed the ICMJE Form for Disclosure of Potential Conflicts of Interest and none were reported.
References
1. Raymond E, Faivre S, Woynarowski JM. et al. Oxaliplatin: mechanism of action and antineoplastic activity. SeminOncol. 1998(Suppl 5):4-12
2. Graf N, Ang WH, Zhu G, Myint M, Lippard SJ. Role of endonucleases XPF and XPG in nucleotide excision repair of platinated DNA and cisplatin/oxaliplatin cytotoxicity. Chembiochem. 2011;12(7):1115-23
3. Lenz HJ, Zhang W, Shi MM. et al. ERCC-1 gene expression levels and outcome to FOLFOX chemotherapy in patients enrolled in CONFIRM1 and CONFIRM2. J Clin Oncol. 2008 abstract4131
4. Orlandi A, Di Salvatore M, Bagalà C, Basso M, Strippoli A, Plastino F, Calegari MA, Cassano A, Astone A, Barone C. ERCC1 Induction after Oxaliplatin Exposure May Depend on KRAS Mutational Status in Colorectal Cancer Cell Line: In Vitro Veritas. J Cancer. 2015Jan1;6(1):70-81
5. Seetharam RN, Sood A, Basu-Mallick A, Augenlicht LH, Mariadason JM, Goel S. Oxaliplatin resistance induced by ERCC1 up-regulation is abrogated by siRNA-mediated gene silencing in human colorectal cancer cells. Anticancer Res. 2010Jul;30(7):2531-8
Author contact
![]() Corresponding author: Armando Orlandi MD, PhD, Department of Medical Oncology, Università Cattolica del Sacro Cuore, Roma. Largo Gemelli, 8 - 00168 Rome, Italy. e-mail: armando.orlandirm.unicatt.it; Phone: +39 06 30156318; Fax: +39 06 30154838.
Corresponding author: Armando Orlandi MD, PhD, Department of Medical Oncology, Università Cattolica del Sacro Cuore, Roma. Largo Gemelli, 8 - 00168 Rome, Italy. e-mail: armando.orlandirm.unicatt.it; Phone: +39 06 30156318; Fax: +39 06 30154838.